Regulatory Evolution
Differential gene expression analysis using RNA-seq
 There are still many unanswered questions concerning the computational analysis of RNA-seq:
There are still many unanswered questions concerning the computational analysis of RNA-seq:
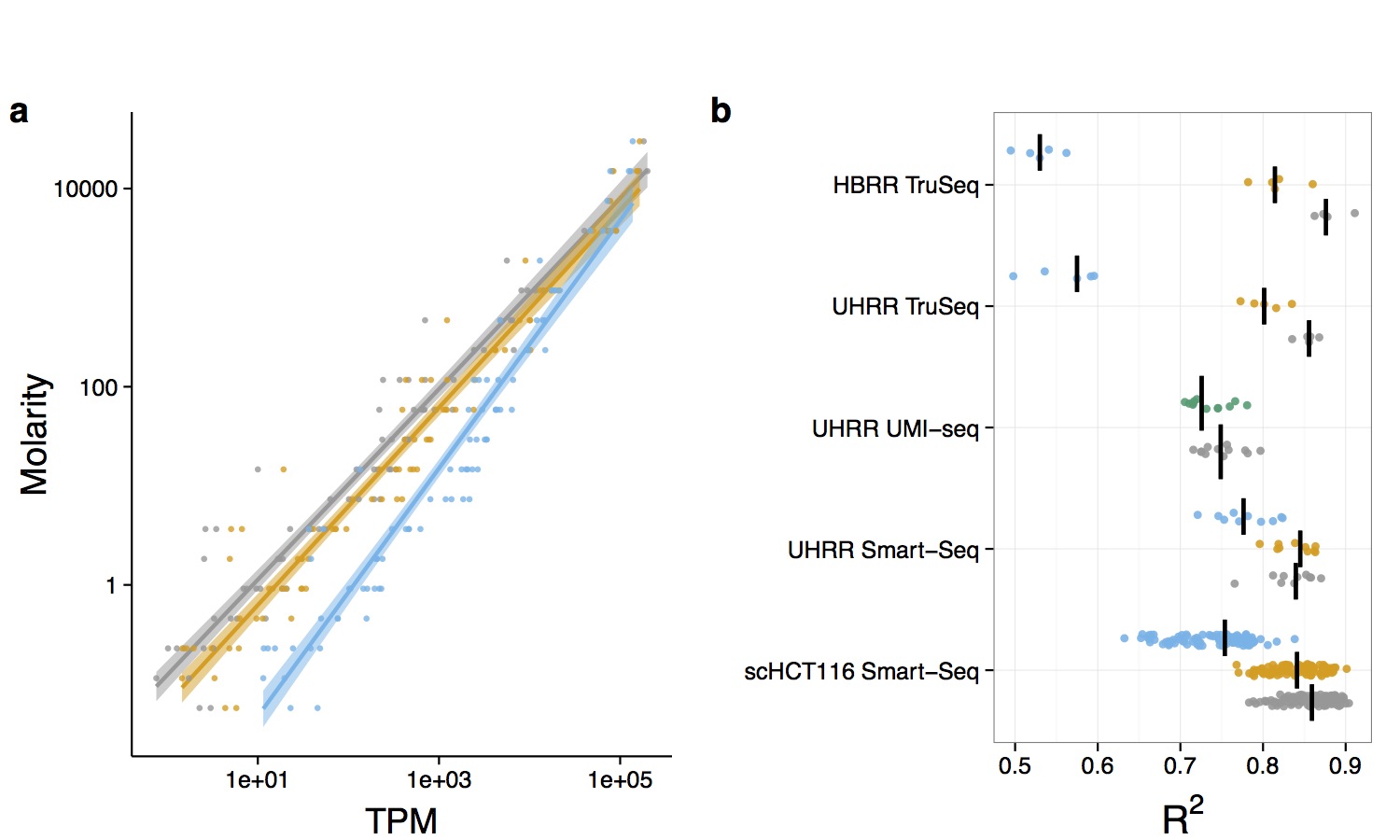
What is the impact of amplification on quantification?
How can I detect DE-genes between species?
Accuracy of RNA-quantification for different
library preparation methods and read-duplicate
treatement strategies.
Modes of evolution in regulatory elements
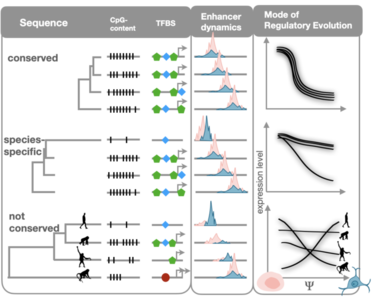
To understand the evolutionary forces that drive promoter and enhancer evolution, we device population genetic and phylogenetic measures of selection in proximal and distal regulatory elements. Using this data we want to identify the genomic factors that drive the measured selection, such as the CpG-island character of the regulatory element and the expression profile of associated genes (Orchard et al. in prep).
The dynamics between epigenetic modifications, mutation and selection
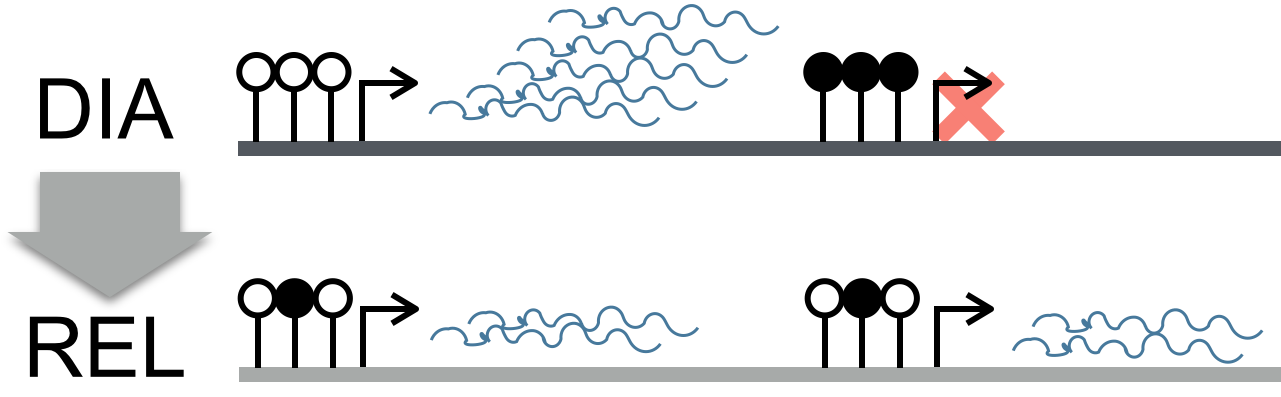 DNA-methylation provides a natural means to link functional changes and mutation rates. The hypermethylation of a CpG-Island promoter silences transcription and at the same time increases the mutation rate of methylated Cs. From an evolutionary perspective, this makes gene regulation in germline tissues via DNA methylation particularly interesting.
DNA-methylation provides a natural means to link functional changes and mutation rates. The hypermethylation of a CpG-Island promoter silences transcription and at the same time increases the mutation rate of methylated Cs. From an evolutionary perspective, this makes gene regulation in germline tissues via DNA methylation particularly interesting.

