mcSCRB-seq
Single-cell RNA sequencing (scRNA-seq) has emerged as a central genome-wide method to characterize cellular identities and processes. Consequently, improving its sensitivity, flexibility and cost-efficiency can advance many research questions. Among the flexible plate-based methods, “Single-Cell RNA-Barcoding and Sequencing” (SCRB-seq) is one of the most sensitive and efficient ones. Here, we systematically evaluated experimental conditions of this protocol and find that adding polyethylene glycol considerably increases sensitivity by enhancing cDNA synthesis. Furthermore, using Terra polymerase increases efficiency due to a more even cDNA amplification that requires less sequencing of libraries. We combined these and other improvements to a new scRNA-seq library protocol we call “molecular crowding SCRB-seq” (mcSCRB-seq), which we show to be the most sensitive and one of the most efficient and flexible scRNA-seq methods to date.
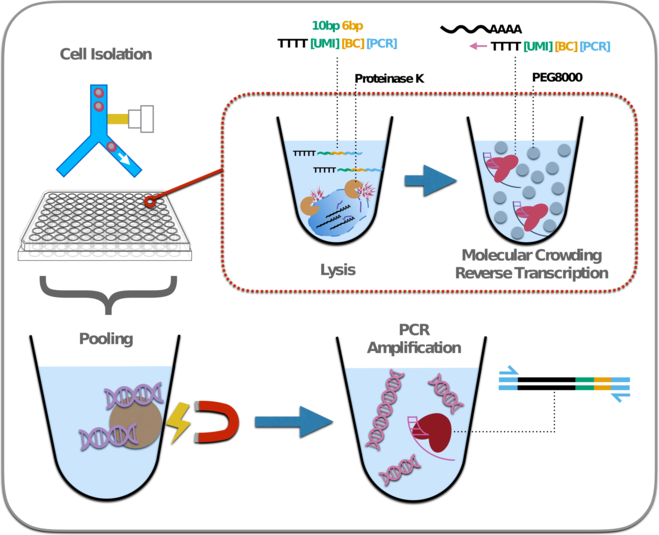
We have used mcSCRB-seq for numerous scRNA-seq projects including complex experimental designs rare cell types and up to 5760 single cells (60 x 96 well plates) in a single experiment. Even though the throughput is lower in comparison to droplet based techniques, this method is very powerful when many conditions are being compared across replicates or the cell type of interest makes up only a small fraction of the cell population and an enrichment using FACS is necessary.
Links:
Paper : https://www.nature.com/articles/s41467-018-05347-6
protocols.io link: https://www.protocols.io/view/mcscrb-seq-protocol-p9kdr4w
Slack: https://publicslack.com/slacks/mcscrb-seq/invites/new

